NOTE: all figures in this post were made by the author using
, LATEX numpy, andmatplotlib
We use distance formulas in
Euclidean Distance
The first—and most common—distance formula is the Euclidean distance.
This is calculated by finding the difference between elements in list
Euclidean distance is a straightforward measure of spatial similarity, making it suitable for many applications. It is used when the features have a clear geometric interpretation, and scale differences between features are not a concern as scale difference is a major drawback with this metric.
Manhattan Distance
This distance formula is different from Euclidean distance because it does not measure the magnitude nor the angle of the line connecting two points. In certain instances, knowing the magnitude of the line between two points is necessary in a
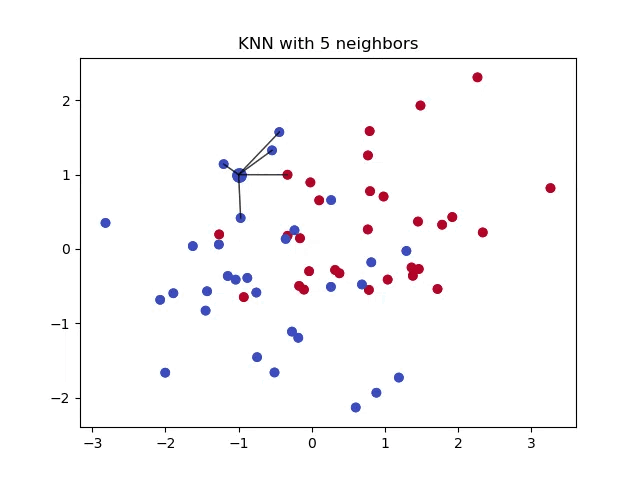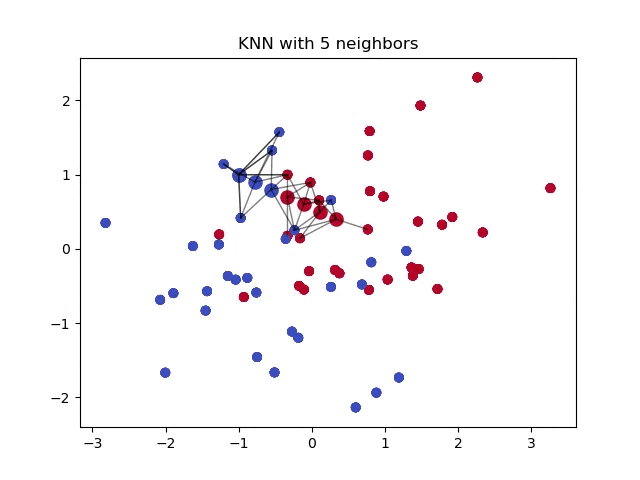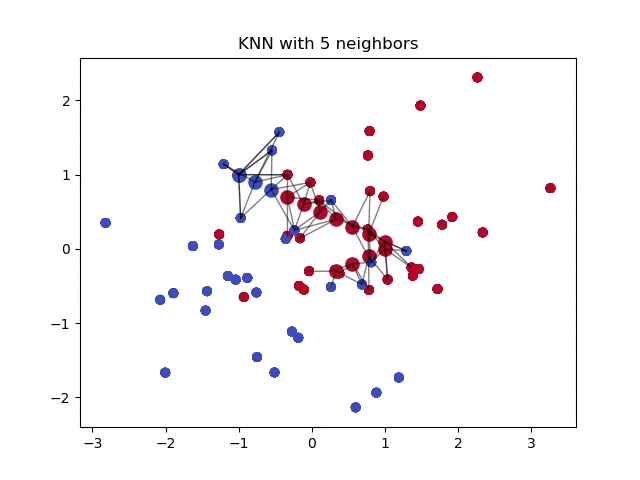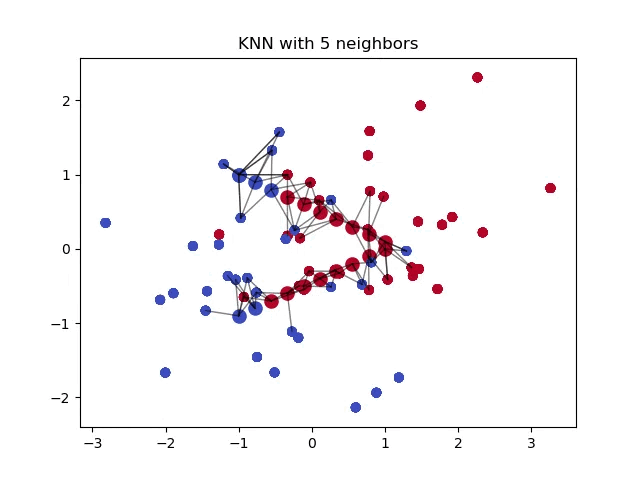
When classifying a point, a shorter distance between that point and another point of a different class often indicates a higher similarity between the points. Consequently, the point is more likely to belong to the class that is closer to it.
You can see the difference in Euclidean distance and Manhattan distance more clearly in the image below. The formula on the right resembles the distance from one street to another in a city grid, hence the name “Manhattan” distance.
The Manhattan distance can be particularly useful in datasets or scenarios where the features have different units of measurement that are all independent of each other. It captures the total discrepancy along each feature dimension without assuming any specific relationship between them.
When calculating the similarity or distance between two houses, using the Euclidean distance would implicitly assume that the features contribute equally to the overall similarity with a straight line connecting them. However, in reality, the differences in square footage, distance to a local school, number of bedrooms, etc. might not have equal importance.
Minkowski Distance
This distance formula is unique in that it includes both Euclidean and Manhattan distances as special cases, when
Note that sklearn.neighbors.KNeighborsClassifier() function uses Minkowski distance as the default metric, most likely because of its versatility. Refer to scipy.spatial.distance() for a complete list of distance metrics.
In general, a higher value of
Cosine Similarity
If you’ve taken a linear algebra class, you’ve definitely seen this formula before. This equation calculates
In a linear algebra textbook, you might see a similar equation that looks like this:
This is the same formula, where
In the example above, the cosine similarity between the two vectors is
You can see the differences in each of the three cosine similarities below. In the leftmost graph, since the two vectors are perpendicular, they have no similarity.
In the middle graph, since the two vectors are multiples of each other, and they have the same direction, they fall on the same line in 2D space. This means they are essentially the same vector—exact similarity—just with a different magnitude. This is apparent because we have
Finally, in the rightmost graph, these two vectors are exactly dissimilar, with a similarity of
Hamming Distance
If we wanted to classify a binary output, this is the metric we want to use. The function